Directional selection
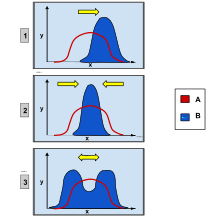
In population genetics, directional selection is a type of natural selection in which one extreme phenotype is favored over both the other extreme and moderate phenotypes. This genetic selection causes the allele frequency to shift toward the chosen extreme over time as allele ratios change from generation to generation. The advantageous extreme allele will increase in frequency among the population as a consequence of survival and reproduction differences among the different present phenotypes in the population. The allele fluctuations as a result of directional selection can be independent of the dominance of the allele, and in some cases if the allele is recessive, it can eventually become fixed in the population.[1][2]
Directional selection was first identified and described by naturalist Charles Darwin in his book On the Origin of Species published in 1859.[3] He identified it as a type of natural selection along with stabilizing selection and disruptive selection.[4] These types of selection also operate by favoring a specific allele and influencing the population's future phenotypic ratio. Disruptive selection favors both extreme phenotypes while the moderate phenotype will be selected against. The frequency of both extreme alleles will increase while the frequency of the moderate allele will decrease, differing from the trend in directional selection in which only one extreme allele is favored. Stabilizing selection favors the moderate phenotype and will select against both extreme phenotypes.[5] Directional selection can be observed in finch beak size, peppered moth color, African cichlid mouth types, and sockeye salmon migration periods.
If there is continuous allele frequency change as a result of directional selection generation from generation, there will be observable changes in the phenotypes of the entire population over time. Directional selection can change the genotypic and phenotypic variation of a population and cause a trend toward one specific phenotype.[6] This selection is an important mechanism in the selection of complex and diversifying traits, and is also a primary force of speciation.[7] Changes in a genotype and consequently a phenotype can either be advantageous, harmful, or neutral and depend on the environment in which the phenotypic shift is happening.[8]
Evidence
[edit]Detection methods
[edit]Directional selection most often occurs during environmental changes or population migrations to new areas with different environmental pressures. Directional selection allows for swift changes in allele frequency that can accompany rapidly changing environmental factors and plays a major role in speciation.[7] Analysis on quantitative trait locus (QTL) effects has been used to examine the impact of directional selection in phenotypic diversification. QTL is a region of a gene that corresponds to a specific phenotypic trait, and the measuring the statistical frequencies of the traits can be helpful in analyzing phenotypic trends.[9] In one study, the analysis showed that directional changes in QTLs affecting various traits were more common than expected by chance among diverse species. This was an indication that directional selection is a primary cause of the phenotypic diversification that can eventually result in speciation.[10]
There are different statistical tests that can be run to test for the presence of directional selection in a population. A highly indicative test of changes in allele frequencies is the QTL sign test, and other tests include the Ka/Ks ratio test and the relative rate test. The QTL sign test compares the number of antagonistic QTL to a neutral model, and allows for testing of directional selection against genetic drift.[11] The Ka/Ks ratio test compares the number of non-synonymous to synonymous substitutions, and a ratio that is greater than 1 indicates directional selection.[12] The relative ratio test looks at the accumulation of advantageous traits against a neutral model, but needs a phylogenetic tree for comparison. This can prove difficult if the full phylogenic history is not known or is not specific enough for the test comparison.[13]
Examples
[edit]Finch beak size
[edit]
Another example of directional selection is the beak size in a specific population of finches. Darwin first observed this in the publication of his book, On the Origin of Species, and he details how the size of the finches beak differs based on environmental factors. On the Galápagos Islands west of the coast of Ecuador, there were groups of finches displaying different beak phenotypes.[14] In one group, the beaks ranged from large and tough to small and smooth. Throughout the wet years, small seeds were more common than large seeds, and because of the large supply of small seeds the finches rarely ate large seeds. During the dry years, neither the small or large seeds were in great abundance, and the birds trended towards eating larger seeds. The changes in diet of the finches based on the environmental wet and dry seasons affected the depth of the birds’ beaks in future generations. [15] The beaks most beneficial to the more plentiful type of seed would be selected for because the birds were able to feed themselves and reproduce.
Peppered moths
[edit]

A significant example of directional selection in populations is the fluctuations of light and dark phenotypes in peppered moths in the 1800s.[16] During the industrial revolution, environmental conditions were rapidly changing with the newfound emission of dark, black smoke from factories that would change the color of trees, rocks, and other niches of moths.[17] Before the industrial revolution, the most prominent phenotype in the peppered moth population was the lighter, speckled moths. They thrived on the light birch trees and their phenotype would provide them with better camouflage from predators. After the Industrial Revolution as the trees become darker with soot, the moths with the darker phenotype were able to blend in and avoid predators better than their white counterparts. As time went on, the darker moths were positively directionally selected for and the allele frequency began to shift due to the increase in the number of darker moths.[18]
African cichlids
[edit]

African cichlids are known to be a diverse fish species, with evidence indicating that they evolved extremely quickly. These fish evolved within the same habitat, but have a variety of morphologies, especially pertaining to the mouth and jaw. Experiments pertaining the cichlid jaw phenotypes was done by Albertson and others in 2003 by crossing two species of African cichlids with very different mouth morphologies. The cross between Labeotropheus fuelleborni (subterminal mouth for biting algae off rocks) and Metriaclima zebra (terminal mouth for suction feeding) allowed for mapping of QTLs affecting feeding morphology. Using the QTL sign test, definitive evidence was used to support the existence of directional selection in the oral jaw apparatus in African cichlids. However, this was not the case for the suspensorium or skull QTLs, suggesting genetic drift or stabilizing selection as mechanisms for the speciation.[19]
Sockeye salmon
[edit]
Sockeye salmon are one of the many species of fish that are anadromous, in which individuals migrate to the same rivers in which they were born to reproduce. These migrations happen around the same time every year, but a 2007 study shows that sockeye salmon found in the waters of the Bristol Bay in Alaska have recently undergone directional selection on the timing of migration.[20] In this study, two populations of sockeye salmon, Egegik and Ugashik, were observed. Data from 1969–2003 provided by the Alaska Department of Fish and Game were divided into five sets of seven years and plotted for average arrival to the fishery. After analyzing the data, it was determined that in both populations the average migration date was earlier and the populations were undergoing directional selection as a result of changing ecological conditions. The Egegik population experienced stronger selection and the migration date shifted four days. The paper suggests that fisheries can be a factor driving this selection because fishing occurs more often in the later periods of migration (especially in the Egegik district), preventing those fish from reproducing.[21] This discovery also goes to show that in addition to environmental changes, human behaviors can also have massive effects on the selection of species around them.[22]
Bears hunting sockeye salmon
[edit]
Studies carried out in Little Togiak Lake in Alaska, indicate that bear predation has a significant impact on sockeye salmon populations, especially in shallow streams. Bears often focus on larger male salmon and tend to prefer those that have just arrived at the spawning grounds, particularly in smaller streams where they can catch them more easily. This predation may accelerate the aging of salmon by favoring later arrivals. Additionally, the impact of predation varies among different salmon populations based on their habitat and density; it tends to be more selective in areas where fish are readily accessible. While high levels of bear predation can occur, healthy salmon populations usually maintain strong reproductive potential, although the effects are more pronounced when populations are low. Overall, these dynamics illustrate how bear predation affects salmon behavior and life cycles, influencing their evolutionary processes. [23]
Large Felids
[edit]This study examines the role of lineage-specific directional selection on body size evolution in felids, revealing that several species, including those in the Panthera genus (lions, tigers, leopards, jaguars, snow leopards), the cheetah, and the puma, exhibit evidence of directional selection favoring larger body mass. These larger body sizes are likely linked to hunting large prey and solitary hunting strategies, which favor physical strength and size. Conversely, the clouded leopard did not show evidence of directional selection for body size, suggesting different ecological pressures, and the jaguarundi showed no clear selection for smaller size despite being smaller than its relatives. These findings highlight that body size evolution in felids is not uniform and is strongly influenced by ecological factors such as prey size and hunting behavior. The study concludes that directional selection for increased body size is likely associated with the need for larger predators to capture large prey, and solitary hunting may accelerate this selection, although the evolutionary paths for different felid lineages can vary considerably.[24]
Soapberry Bugs
[edit]
Soapberry Bugs (Jadera haematoloma) primarily feed on seeds produced by plants of the Sapindaceae family. These soapberry bugs use their beaks to feed on the seeds within the fruits of these plants, so it is crucial that their beak size is long enough to reach the seeds from the exterior of the fruits. However, the distance from the exterior of the fruit to the seed can vary. Scott Carroll and Christin Boyd (1992) conducted an experiment where they would observe how three newly introduced plant species introduced to North America that were colonized by these soapberry bugs would affect the natural selection of the insect’s beak length. Each new plant species hosted fruits of different sizes compared to the native hosts. They found that there was indeed a close correlation between the radius of the fruit and the length of the beak. There was a positive directional selection for larger beaks when the radius of the fruit was larger, and there was a positive directional selection for smaller beaks when the radius of the fruit was smaller. To confirm that these differences were caused by genetic differences and not through phenotypic plasticity, Carroll raised young soapberry bugs from the populations based on the introduced plant species and found that their beak length was retained when they were developed on the alternative host. [25]
Ecological impact
[edit]Directional selection can quickly lead to vast changes in allele frequencies in a population because of the cumulative nature of reproduction of the fittest. Because the main cause for directional selection is different and changing environmental pressures, rapidly changing environments, such as those affected by climate change, can cause drastic changes within populations.
Diversity
[edit]Limiting the number of genotypes in a certain population can be deleterious to the ecosystem as a whole by shrink the potential genetic gene pool.[26] Low amount of genetic variation can lead to mass extinctions and endangered species because of the large impact one mutation can have on the entire population if there are only a few specific genes present throughout.
Urban Influence
[edit]It is important to note the impact that humans have on genetic diversity as well, and be aware of the ways to reduce harmful impacts on natural environments.[27] Major roads, waterway pollution, and urbanization all cause environmental selection and could potentially result in changes in allele frequencies.[28] Hunting may also play a role in directional selection, albeit more so in smaller populations.
Timescale
[edit]Typically directional selection acts strongly for short bursts and is not sustained over long periods of time.[29] If it was sustained, a population might hit biological constraints such that it no longer responds to selection. However, it is possible for directional selection to take a very long time to find a local optimum on a fitness landscape.[30] A possible example of long-term directional selection is the tendency of proteins to become more hydrophobic over time,[31] and to have their hydrophobic amino acids more interspersed along the sequence.[32]
See also
[edit]- Adaptive evolution in the human genome
- Balancing selection
- Disruptive selection
- Frequency-dependent foraging by pollinators
- Negative selection (natural selection)
- Stabilizing selection
- Peppered moth evolution
- Fluctuating selection
References
[edit]- ^ Molles, MC (2010). Ecology Concepts and Applications. McGraw-Hill Higher Learning.
- ^ Teshima, Kosuke M.; Przeworski, Molly (January 2006). "Directional Positive Selection on an Allele of Arbitrary Dominance". Genetics. 172 (1): 713–718. doi:10.1534/genetics.105.044065. PMC 1456198. PMID 16219788.
- ^ Kaiser, Margaret (November 2014). "First editions of Darwin's 'Origin of Species'". National Library of Medicine.
- ^ Darwin, C (1859). On the origin of species by means of natural selection, or the preservation of favoured races in the struggle for life. London: John Murray.
- ^ Mitchell-Olds, Thomas; Willis, John H.; Goldstein, David B. (2007). "Which evolutionary processes influence natural genetic variation for phenotypic traits?". Nature Reviews Genetics. 8 (11). Springer Nature: 845–856. doi:10.1038/nrg2207. ISSN 1471-0056. PMID 17943192. S2CID 14914998.
- ^ Melo, Diogo; Marroig, Gabriel (January 2015). "Directional selection can drive the evolution of modularity in complex traits". Proceedings of the National Academy of Sciences of the United States of America. 112 (2): 470–475. Bibcode:2015PNAS..112..470M. doi:10.1073/pnas.1322632112. PMC 4299217. PMID 25548154.
- ^ a b Rieseberg, Loren H.; Widmer, Alex; Arntz, A. Michele; Burke, John M. (September 2002). "Directional selection is the primary cause of phenotypic diversification". Proceedings of the National Academy of Sciences. 99 (19): 12242–12245. Bibcode:2002PNAS...9912242R. doi:10.1073/pnas.192360899. PMC 129429. PMID 12221290.
- ^ Thiltgen, Grant; dos Reis, Mario; Goldstein, Richard A. (December 2016). "Finding Direction in the Search for Selection". Journal of Molecular Evolution. 84 (1): 39–50. doi:10.1007/s00239-016-9765-5. PMC 5253163. PMID 27913840.
- ^ Powder, Kara E. (March 2024). "Quantitative Trait Loci (QTL) Mapping". EQTL Analysis. Methods in Molecular Biology. Vol. 2082. pp. 211–229. doi:10.1007/978-1-0716-0026-9_15. ISBN 978-1-0716-0025-2. PMID 31849018.
- ^ Rieseberg, Loren H.; Widmer, Alex; Arntz, A. Michele; Burke, John M. (2002-09-17). "Directional selection is the primary cause of phenotypic diversification". Proceedings of the National Academy of Sciences of the United States of America. 99 (19): 12242–5. Bibcode:2002PNAS...9912242R. doi:10.1073/pnas.192360899. PMC 129429. PMID 12221290.
- ^ Orr, H.A. (1998). "Testing Natural Selection vs. Genetic Drift in Phenotypic Evolution Using Quantitative Trait Locus Data". Genetics. 149 (4): 2099–2104. doi:10.1093/genetics/149.4.2099. PMC 1460271. PMID 9691061.
- ^ Hurst, Laurence D (2002). "The Ka/Ks ratio: diagnosing the form of sequence evolution". Trends in Genetics. 18 (9). Elsevier BV: 486–487. doi:10.1016/s0168-9525(02)02722-1. ISSN 0168-9525. PMID 12175810.
- ^ Creevey, Christopher J.; McInerney, James O. (2002). "An algorithm for detecting directional and non-directional positive selection, neutrality and negative selection in protein coding DNA sequences". Gene. 300 (1–2). Elsevier BV: 43–51. doi:10.1016/s0378-1119(02)01039-9. ISSN 0378-1119. PMID 12468084.
- ^ Burrows, Leah (November 2021). "For Darwin's finches, beak shape goes beyond evolution". Harvard School of Engineering.
- ^ Campbell, Neil A.; Reece, Jane B. (2002). Biology (6th ed.). Benjamin Cummings. pp. 450–451. ISBN 978-0-8053-6624-2.
- ^ "Peppered Moth". globalchange.umich.edu. Retrieved 2024-03-24.
- ^ "Peppered Moth and natural selection". butterfly-conservation.org. Retrieved 2024-03-24.
- ^ Saccheri, Ilik J. (October 2008). "Selection and gene flow on a diminishing cline of melanic peppered moths". Proceedings of the National Academy of Sciences. 105 (42): 16212–16217. Bibcode:2008PNAS..10516212S. doi:10.1073/pnas.0803785105. PMC 2571026. PMID 18854412.
- ^ Albertson, R. C.; Streelman, J. T.; Kocher, T. D. (2003-04-18). "Directional selection has shaped the oral jaws of Lake Malawi cichlid fishes". Proceedings of the National Academy of Sciences. 100 (9): 5252–5257. Bibcode:2003PNAS..100.5252A. doi:10.1073/pnas.0930235100. ISSN 0027-8424. PMC 154331. PMID 12704237.
- ^ Quinn, Thomas P. (April 2007). "Directional selection by fisheries and the timing of sockeye salmon (Oncorhynchus Nerka) Migrations". Ecological Applications. 17 (3): 731–739. Bibcode:2007EcoAp..17..731Q. doi:10.1890/06-0771. PMID 17494392.
- ^ Quinn, Thomas P.; Hodgson, Sayre; Flynn, Lucy; Hilborn, Ray; Rogers, Donald E. (2007). "Directional Selection by Fisheries and the Timing of Sockeye Salmon (Oncorhynchus Nerka) Migrations". Ecological Applications. 17 (3). Wiley: 731–739. Bibcode:2007EcoAp..17..731Q. doi:10.1890/06-0771. ISSN 1051-0761. PMID 17494392.
- ^ Lin, J. E.; Hard, J. J.; Naish, K. A.; Peterson, D.; Hilborn, R.; Hauser, L. (May 2016). "It's a bear market: evolutionary and ecological effects of predation on two wild sockeye salmon populations". Heredity. 116 (5): 447–457. doi:10.1038/hdy.2016.3. ISSN 1365-2540. PMC 4834386.
- ^ Lin, J. E.; Hard, J. J.; Naish, K. A.; Peterson, D.; Hilborn, R.; Hauser, L. (May 2016). "It's a bear market: evolutionary and ecological effects of predation on two wild sockeye salmon populations". Heredity. 116 (5): 447–457. doi:10.1038/hdy.2016.3. ISSN 1365-2540. PMC 4834386.
- ^ Harano, Tomohiro; Kutsukake, Nobuyuki (2023-03-01). "Way to big cats: Directional selection in body size evolution in living felids". Journal of Mammalian Evolution. 30 (1): 97–108. doi:10.1007/s10914-022-09639-z. ISSN 1573-7055.
- ^ Carroll, Scott P.; Boyd, Christin (1 August 1992). "HOST RACE RADIATION IN THE SOAPBERRY BUG: NATURAL HISTORY WITH THE HISTORY". Evolution. 46 (4): 1052–1069. doi:10.1111/j.1558-5646.1992.tb00619.x. ISSN 0014-3820.
- ^ Star, Bastiaan; Spencer, Hamish G. (May 2013). "Effects of Genetic Drift and Gene Flow on the Selective Maintenance of Genetic Variation". Genetics. 194 (1): 235–244. doi:10.1534/genetics.113.149781. PMC 3632471. PMID 23457235.
- ^ Mysterud, Atle (13 May 2011). "Selective harvesting of large mammals: how often does it result in directional selection?". Journal of Applied Ecology. 48 (4): 827–834. doi:10.1111/j.1365-2664.2011.02006.x.
- ^ Hunter, Philip (April 2007). "The human impact on biological diversity". EMBO Reports. 8 (4): 316–318. doi:10.1038/sj.embor.7400951. PMC 1852758. PMID 17401404.
- ^ Hoekstra, H. E.; Hoekstra, J. M.; Berrigan, D.; Vignieri, S. N.; Hoang, A.; Hill, C. E.; Beerli, P.; Kingsolver, J. G. (2001-07-24). "Strength and tempo of directional selection in the wild". Proceedings of the National Academy of Sciences. 98 (16): 9157–9160. Bibcode:2001PNAS...98.9157H. doi:10.1073/pnas.161281098. ISSN 0027-8424. PMC 55389. PMID 11470913.
- ^ Kaznatcheev, Artem (May 2019). "Computational Complexity as an Ultimate Constraint on Evolution". Genetics. 212 (1): 245–265. doi:10.1534/genetics.119.302000. PMC 6499524. PMID 30833289.
- ^ Wilson, Benjamin A.; Foy, Scott G.; Neme, Rafik; Masel, Joanna (24 April 2017). "Young genes are highly disordered as predicted by the preadaptation hypothesis of de novo gene birth" (PDF). Nature Ecology & Evolution. 1 (6): 0146–146. Bibcode:2017NatEE...1..146W. doi:10.1038/s41559-017-0146. hdl:10150/627822. PMC 5476217. PMID 28642936.
- ^ Foy, Scott G.; Wilson, Benjamin A.; Bertram, Jason; Cordes, Matthew H. J.; Masel, Joanna (April 2019). "A Shift in Aggregation Avoidance Strategy Marks a Long-Term Direction to Protein Evolution". Genetics. 211 (4): 1345–1355. doi:10.1534/genetics.118.301719. PMC 6456324. PMID 30692195.
Further reading
[edit]- Sabeti PC; et al. (2006). "Positive Natural Selection in the Human Lineage". Science. 312 (5780): 1614–1620. Bibcode:2006Sci...312.1614S. doi:10.1126/science.1124309. PMID 16778047. S2CID 10809290.
- Pickrell JK, Coop G, Novembre J, et al. (May 2009). "Signals of recent positive selection in a worldwide sample of human populations". Genome Research. 19 (5): 826–837. doi:10.1101/gr.087577.108. PMC 2675971. PMID 19307593.
- Types of Selection
- Natural Selection
- Modern Theories of Evolution
